This page is part of the FHIR Specification (v5.0.0-ballot: FHIR R5 Ballot Preview). The current version which supercedes this version is 5.0.0. For a full list of available versions, see the Directory of published versions  . Page versions: R5 R4B R4
. Page versions: R5 R4B R4
Clinical Genomics  Work Group Work Group | Maturity Level: 1 | Trial Use | Security Category: Patient | Compartments: Patient |
Representation of a molecular sequence.
The MolecularSequence resource is designed for representing molecular sequences. It can represent the sequence in different ways, allowing implementations to adopt the most effective one for their use case.
It is strongly encouraged to provide as much information in this resource for any reported sequences, because receiving systems (e.g. discovery research, outcomes analysis, and public health reporting) may use this information to normalize sequences over time or across sources. However, these data should not be used to dynamically correct/change sequence representations for clinical use outside of the laboratory, due to insufficient information.
The MolecularSequence resource is designed to represent a single sequence in an instance. Each sequence might have multiple representations, but implementers SHALL ensure all representations are for the same sequence. This means that if a single MolecularSequence instance contains a literal, two formatted files, and a relative, all four of those representions must represent the same sequence. This can be a challenge across systems, as semantic equivalency of sequences cannot be guaranteed unless there is an agreed upon standard between sending and receiving systems.
The MolecularSequence resource should only be used to capture a molecular sequence. It will not be used for other entities such as variant, variant annotations, genotypes, haplotypes, etc. Those concepts will be captured in Observation profiles found in the Genomics Reporting Implementation Guide  . The sequence that was observed that led to the identification of those concepts can be delivered with this resource, and will be referenced by those observations.
. The sequence that was observed that led to the identification of those concepts can be delivered with this resource, and will be referenced by those observations.
MolecularSequence will not be used to capture data such as precise read of DNA sequences and sequence alignment are not included; such data may be accessible through references to GA4GH  (Global Alliance for Genomics and Health) API, and may be referenced to by the
(Global Alliance for Genomics and Health) API, and may be referenced to by the formatted element.
Structure
| Name | Flags | Card. | Type | Description & Constraints |
|---|---|---|---|---|
 | TU | DomainResource | Representation of a molecular sequence Elements defined in Ancestors: id, meta, implicitRules, language, text, contained, extension, modifierExtension | |
  | Σ | 0..* | Identifier | Unique ID for this particular sequence |
  | Σ | 0..1 | code | aa | dna | rna sequenceType (Required) |
  | Σ | 0..1 | Reference(Patient | Group | Device | Location | Organization | Procedure | Practitioner | Medication | Substance | BiologicallyDerivedProduct | NutritionProduct) | Subject this sequence is associated too |
  | Σ | 0..1 | Reference(Specimen) | Specimen used for sequencing |
  | Σ | 0..1 | Reference(Device) | The method for sequencing |
  | Σ | 0..1 | Reference(Organization) | Who should be responsible for test result |
  | Σ | 0..1 | string | Sequence that was observed |
  | Σ | 0..* | Attachment | Embedded file or a link (URL) which contains content to represent the sequence |
  | Σ | 0..* | BackboneElement | A sequence defined relative to another sequence |
   | Σ | 1..1 | CodeableConcept | Ways of identifying nucleotides or amino acids within a sequence LL5323-2  (Extensible) (Extensible) |
   | 0..1 | integer | Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together | |
   | 0..1 | Range | Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together | |
   | ΣC | 0..1 | BackboneElement | A sequence used as starting sequence + Rule: Both genomeAssembly and chromosome must be both contained if either one of them is contained + Rule: Have and only have one of the following elements in startingSequence: 1. genomeAssembly; 2 sequence |
    | Σ | 0..1 | CodeableConcept | The genome assembly used for starting sequence, e.g. GRCh38 LL1040-6  (Extensible) (Extensible) |
    | Σ | 0..1 | CodeableConcept | Chromosome Identifier LL2938-0  (Required) (Required) |
    | Σ | 0..1 | The reference sequence that represents the starting sequence Multiple bindings acceptable (NCBI or LRG) (Example) | |
     | CodeableConcept | |||
     | string | |||
     | Reference(MolecularSequence) | |||
    | Σ | 0..1 | integer | Start position of the window on the starting sequence |
    | Σ | 0..1 | integer | End position of the window on the starting sequence |
    | Σ | 0..1 | code | sense | antisense orientationType (Required) |
    | Σ | 0..1 | code | watson | crick strandType (Required) |
   | Σ | 0..* | BackboneElement | Changes in sequence from the starting sequence |
    | Σ | 0..1 | integer | Start position of the edit on the starting sequence |
    | Σ | 0..1 | integer | End position of the edit on the starting sequence |
    | Σ | 0..1 | string | Allele that was observed |
    | Σ | 0..1 | string | Allele in the starting sequence |
 Documentation for this format Documentation for this format | ||||
See the Extensions for this resource
UML Diagram (Legend)
XML Template
<MolecularSequence xmlns="http://hl7.org/fhir"><!-- from Resource: id, meta, implicitRules, and language --> <!-- from DomainResource: text, contained, extension, and modifierExtension --> <identifier><!-- 0..* Identifier Unique ID for this particular sequence --></identifier> <type value="[code]"/><!-- 0..1 aa | dna | rna --> <subject><!-- 0..1 Reference(BiologicallyDerivedProduct|Device|Group|Location| Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure| Substance) Subject this sequence is associated too --></subject> <specimen><!-- 0..1 Reference(Specimen) Specimen used for sequencing --></specimen> <device><!-- 0..1 Reference(Device) The method for sequencing --></device> <performer><!-- 0..1 Reference(Organization) Who should be responsible for test result --></performer> <literal value="[string]"/><!-- 0..1 Sequence that was observed --> <formatted><!-- 0..* Attachment Embedded file or a link (URL) which contains content to represent the sequence --></formatted> <relative> <!-- 0..* A sequence defined relative to another sequence --> <coordinateSystem><!-- 1..1 CodeableConcept Ways of identifying nucleotides or amino acids within a sequence
--></coordinateSystem> <ordinalPosition value="[integer]"/><!-- 0..1 Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together --> <sequenceRange><!-- 0..1 Range Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together --></sequenceRange> <startingSequence> <!-- 0..1 A sequence used as starting sequence --> <genomeAssembly><!-- 0..1 CodeableConcept The genome assembly used for starting sequence, e.g. GRCh38
--></genomeAssembly> <chromosome><!-- 0..1 CodeableConcept Chromosome Identifier
--></chromosome> <sequence[x]><!-- 0..1 CodeableConcept|string|Reference(MolecularSequence) The reference sequence that represents the starting sequence --></sequence[x]> <windowStart value="[integer]"/><!-- 0..1 Start position of the window on the starting sequence --> <windowEnd value="[integer]"/><!-- 0..1 End position of the window on the starting sequence --> <orientation value="[code]"/><!-- 0..1 sense | antisense --> <strand value="[code]"/><!-- 0..1 watson | crick --> </startingSequence> <edit> <!-- 0..* Changes in sequence from the starting sequence --> <start value="[integer]"/><!-- 0..1 Start position of the edit on the starting sequence --> <end value="[integer]"/><!-- 0..1 End position of the edit on the starting sequence --> <replacementSequence value="[string]"/><!-- 0..1 Allele that was observed --> <replacedSequence value="[string]"/><!-- 0..1 Allele in the starting sequence --> </edit> </relative> </MolecularSequence>
JSON Template
{ "resourceType" : "MolecularSequence",
// from Resource: id, meta, implicitRules, and language
// from DomainResource: text, contained, extension, and modifierExtension
"identifier" : [{ Identifier }], // Unique ID for this particular sequence
"type" : "<code>", // aa | dna | rna
"subject" : { Reference(BiologicallyDerivedProduct|Device|Group|Location|
Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure|
Substance) }, // Subject this sequence is associated too
"specimen" : { Reference(Specimen) }, // Specimen used for sequencing
"device" : { Reference(Device) }, // The method for sequencing
"performer" : { Reference(Organization) }, // Who should be responsible for test result
"literal" : "<string>", // Sequence that was observed
"formatted" : [{ Attachment }], // Embedded file or a link (URL) which contains content to represent the sequence
"relative" : [{ // A sequence defined relative to another sequence
"coordinateSystem" : { CodeableConcept }, // R! Ways of identifying nucleotides or amino acids within a sequence
"resourceType" : "MolecularSequence",
// from Resource: id, meta, implicitRules, and language
// from DomainResource: text, contained, extension, and modifierExtension
"identifier" : [{ Identifier }], // Unique ID for this particular sequence
"type" : "<code>", // aa | dna | rna
"subject" : { Reference(BiologicallyDerivedProduct|Device|Group|Location|
Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure|
Substance) }, // Subject this sequence is associated too
"specimen" : { Reference(Specimen) }, // Specimen used for sequencing
"device" : { Reference(Device) }, // The method for sequencing
"performer" : { Reference(Organization) }, // Who should be responsible for test result
"literal" : "<string>", // Sequence that was observed
"formatted" : [{ Attachment }], // Embedded file or a link (URL) which contains content to represent the sequence
"relative" : [{ // A sequence defined relative to another sequence
"coordinateSystem" : { CodeableConcept }, // R! Ways of identifying nucleotides or amino acids within a sequence  "ordinalPosition" : <integer>, // Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together
"sequenceRange" : { Range }, // Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together
"startingSequence" : { // A sequence used as starting sequence
"genomeAssembly" : { CodeableConcept }, // The genome assembly used for starting sequence, e.g. GRCh38
"ordinalPosition" : <integer>, // Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together
"sequenceRange" : { Range }, // Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together
"startingSequence" : { // A sequence used as starting sequence
"genomeAssembly" : { CodeableConcept }, // The genome assembly used for starting sequence, e.g. GRCh38  "chromosome" : { CodeableConcept }, // Chromosome Identifier
"chromosome" : { CodeableConcept }, // Chromosome Identifier  // sequence[x]: The reference sequence that represents the starting sequence. One of these 3:
"sequenceCodeableConcept" : { CodeableConcept },
"sequenceString" : "<string>",
"sequenceReference" : { Reference(MolecularSequence) },
"windowStart" : <integer>, // Start position of the window on the starting sequence
"windowEnd" : <integer>, // End position of the window on the starting sequence
"orientation" : "<code>", // sense | antisense
"strand" : "<code>" // watson | crick
},
"edit" : [{ // Changes in sequence from the starting sequence
"start" : <integer>, // Start position of the edit on the starting sequence
"end" : <integer>, // End position of the edit on the starting sequence
"replacementSequence" : "<string>", // Allele that was observed
"replacedSequence" : "<string>" // Allele in the starting sequence
}]
}]
}
// sequence[x]: The reference sequence that represents the starting sequence. One of these 3:
"sequenceCodeableConcept" : { CodeableConcept },
"sequenceString" : "<string>",
"sequenceReference" : { Reference(MolecularSequence) },
"windowStart" : <integer>, // Start position of the window on the starting sequence
"windowEnd" : <integer>, // End position of the window on the starting sequence
"orientation" : "<code>", // sense | antisense
"strand" : "<code>" // watson | crick
},
"edit" : [{ // Changes in sequence from the starting sequence
"start" : <integer>, // Start position of the edit on the starting sequence
"end" : <integer>, // End position of the edit on the starting sequence
"replacementSequence" : "<string>", // Allele that was observed
"replacedSequence" : "<string>" // Allele in the starting sequence
}]
}]
}
Turtle Template
@prefix fhir: <http://hl7.org/fhir/> .[ a fhir:MolecularSequence; fhir:nodeRole fhir:treeRoot; # if this is the parser root # from Resource: .id, .meta, .implicitRules, and .language # from DomainResource: .text, .contained, .extension, and .modifierExtension fhir:MolecularSequence.identifier [ Identifier ], ... ; # 0..* Unique ID for this particular sequence fhir:MolecularSequence.type [ code ]; # 0..1 aa | dna | rna fhir:MolecularSequence.subject [ Reference(BiologicallyDerivedProduct|Device|Group|Location|Medication|NutritionProduct| Organization|Patient|Practitioner|Procedure|Substance) ]; # 0..1 Subject this sequence is associated too fhir:MolecularSequence.specimen [ Reference(Specimen) ]; # 0..1 Specimen used for sequencing fhir:MolecularSequence.device [ Reference(Device) ]; # 0..1 The method for sequencing fhir:MolecularSequence.performer [ Reference(Organization) ]; # 0..1 Who should be responsible for test result fhir:MolecularSequence.literal [ string ]; # 0..1 Sequence that was observed fhir:MolecularSequence.formatted [ Attachment ], ... ; # 0..* Embedded file or a link (URL) which contains content to represent the sequence fhir:MolecularSequence.relative [ # 0..* A sequence defined relative to another sequence fhir:MolecularSequence.relative.coordinateSystem [ CodeableConcept ]; # 1..1 Ways of identifying nucleotides or amino acids within a sequence fhir:MolecularSequence.relative.ordinalPosition [ integer ]; # 0..1 Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together fhir:MolecularSequence.relative.sequenceRange [ Range ]; # 0..1 Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together fhir:MolecularSequence.relative.startingSequence [ # 0..1 A sequence used as starting sequence fhir:MolecularSequence.relative.startingSequence.genomeAssembly [ CodeableConcept ]; # 0..1 The genome assembly used for starting sequence, e.g. GRCh38 fhir:MolecularSequence.relative.startingSequence.chromosome [ CodeableConcept ]; # 0..1 Chromosome Identifier # MolecularSequence.relative.startingSequence.sequence[x] : 0..1 The reference sequence that represents the starting sequence. One of these 3 fhir:MolecularSequence.relative.startingSequence.sequenceCodeableConcept [ CodeableConcept ] fhir:MolecularSequence.relative.startingSequence.sequenceString [ string ] fhir:MolecularSequence.relative.startingSequence.sequenceReference [ Reference(MolecularSequence) ] fhir:MolecularSequence.relative.startingSequence.windowStart [ integer ]; # 0..1 Start position of the window on the starting sequence fhir:MolecularSequence.relative.startingSequence.windowEnd [ integer ]; # 0..1 End position of the window on the starting sequence fhir:MolecularSequence.relative.startingSequence.orientation [ code ]; # 0..1 sense | antisense fhir:MolecularSequence.relative.startingSequence.strand [ code ]; # 0..1 watson | crick ]; fhir:MolecularSequence.relative.edit [ # 0..* Changes in sequence from the starting sequence fhir:MolecularSequence.relative.edit.start [ integer ]; # 0..1 Start position of the edit on the starting sequence fhir:MolecularSequence.relative.edit.end [ integer ]; # 0..1 End position of the edit on the starting sequence fhir:MolecularSequence.relative.edit.replacementSequence [ string ]; # 0..1 Allele that was observed fhir:MolecularSequence.relative.edit.replacedSequence [ string ]; # 0..1 Allele in the starting sequence ], ...; ], ...; ]
Changes since R4
| MolecularSequence | |
| MolecularSequence.subject |
|
| MolecularSequence.literal |
|
| MolecularSequence.formatted |
|
| MolecularSequence.relative |
|
| MolecularSequence.relative.coordinateSystem |
|
| MolecularSequence.relative.ordinalPosition |
|
| MolecularSequence.relative.sequenceRange |
|
| MolecularSequence.relative.startingSequence |
|
| MolecularSequence.relative.startingSequence.genomeAssembly |
|
| MolecularSequence.relative.startingSequence.chromosome |
|
| MolecularSequence.relative.startingSequence.sequence[x] |
|
| MolecularSequence.relative.startingSequence.windowStart |
|
| MolecularSequence.relative.startingSequence.windowEnd |
|
| MolecularSequence.relative.startingSequence.orientation |
|
| MolecularSequence.relative.startingSequence.strand |
|
| MolecularSequence.relative.edit |
|
| MolecularSequence.relative.edit.start |
|
| MolecularSequence.relative.edit.end |
|
| MolecularSequence.relative.edit.replacementSequence |
|
| MolecularSequence.relative.edit.replacedSequence |
|
| MolecularSequence.coordinateSystem |
|
| MolecularSequence.patient |
|
| MolecularSequence.quantity |
|
| MolecularSequence.referenceSeq |
|
| MolecularSequence.variant |
|
| MolecularSequence.observedSeq |
|
| MolecularSequence.quality |
|
| MolecularSequence.readCoverage |
|
| MolecularSequence.repository |
|
| MolecularSequence.pointer |
|
| MolecularSequence.structureVariant |
|
See the Full Difference for further information
Structure
| Name | Flags | Card. | Type | Description & Constraints |
|---|---|---|---|---|
 | TU | DomainResource | Representation of a molecular sequence Elements defined in Ancestors: id, meta, implicitRules, language, text, contained, extension, modifierExtension | |
  | Σ | 0..* | Identifier | Unique ID for this particular sequence |
  | Σ | 0..1 | code | aa | dna | rna sequenceType (Required) |
  | Σ | 0..1 | Reference(Patient | Group | Device | Location | Organization | Procedure | Practitioner | Medication | Substance | BiologicallyDerivedProduct | NutritionProduct) | Subject this sequence is associated too |
  | Σ | 0..1 | Reference(Specimen) | Specimen used for sequencing |
  | Σ | 0..1 | Reference(Device) | The method for sequencing |
  | Σ | 0..1 | Reference(Organization) | Who should be responsible for test result |
  | Σ | 0..1 | string | Sequence that was observed |
  | Σ | 0..* | Attachment | Embedded file or a link (URL) which contains content to represent the sequence |
  | Σ | 0..* | BackboneElement | A sequence defined relative to another sequence |
   | Σ | 1..1 | CodeableConcept | Ways of identifying nucleotides or amino acids within a sequence LL5323-2  (Extensible) (Extensible) |
   | 0..1 | integer | Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together | |
   | 0..1 | Range | Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together | |
   | ΣC | 0..1 | BackboneElement | A sequence used as starting sequence + Rule: Both genomeAssembly and chromosome must be both contained if either one of them is contained + Rule: Have and only have one of the following elements in startingSequence: 1. genomeAssembly; 2 sequence |
    | Σ | 0..1 | CodeableConcept | The genome assembly used for starting sequence, e.g. GRCh38 LL1040-6  (Extensible) (Extensible) |
    | Σ | 0..1 | CodeableConcept | Chromosome Identifier LL2938-0  (Required) (Required) |
    | Σ | 0..1 | The reference sequence that represents the starting sequence Multiple bindings acceptable (NCBI or LRG) (Example) | |
     | CodeableConcept | |||
     | string | |||
     | Reference(MolecularSequence) | |||
    | Σ | 0..1 | integer | Start position of the window on the starting sequence |
    | Σ | 0..1 | integer | End position of the window on the starting sequence |
    | Σ | 0..1 | code | sense | antisense orientationType (Required) |
    | Σ | 0..1 | code | watson | crick strandType (Required) |
   | Σ | 0..* | BackboneElement | Changes in sequence from the starting sequence |
    | Σ | 0..1 | integer | Start position of the edit on the starting sequence |
    | Σ | 0..1 | integer | End position of the edit on the starting sequence |
    | Σ | 0..1 | string | Allele that was observed |
    | Σ | 0..1 | string | Allele in the starting sequence |
 Documentation for this format Documentation for this format | ||||
See the Extensions for this resource
XML Template
<MolecularSequence xmlns="http://hl7.org/fhir"><!-- from Resource: id, meta, implicitRules, and language --> <!-- from DomainResource: text, contained, extension, and modifierExtension --> <identifier><!-- 0..* Identifier Unique ID for this particular sequence --></identifier> <type value="[code]"/><!-- 0..1 aa | dna | rna --> <subject><!-- 0..1 Reference(BiologicallyDerivedProduct|Device|Group|Location| Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure| Substance) Subject this sequence is associated too --></subject> <specimen><!-- 0..1 Reference(Specimen) Specimen used for sequencing --></specimen> <device><!-- 0..1 Reference(Device) The method for sequencing --></device> <performer><!-- 0..1 Reference(Organization) Who should be responsible for test result --></performer> <literal value="[string]"/><!-- 0..1 Sequence that was observed --> <formatted><!-- 0..* Attachment Embedded file or a link (URL) which contains content to represent the sequence --></formatted> <relative> <!-- 0..* A sequence defined relative to another sequence --> <coordinateSystem><!-- 1..1 CodeableConcept Ways of identifying nucleotides or amino acids within a sequence
--></coordinateSystem> <ordinalPosition value="[integer]"/><!-- 0..1 Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together --> <sequenceRange><!-- 0..1 Range Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together --></sequenceRange> <startingSequence> <!-- 0..1 A sequence used as starting sequence --> <genomeAssembly><!-- 0..1 CodeableConcept The genome assembly used for starting sequence, e.g. GRCh38
--></genomeAssembly> <chromosome><!-- 0..1 CodeableConcept Chromosome Identifier
--></chromosome> <sequence[x]><!-- 0..1 CodeableConcept|string|Reference(MolecularSequence) The reference sequence that represents the starting sequence --></sequence[x]> <windowStart value="[integer]"/><!-- 0..1 Start position of the window on the starting sequence --> <windowEnd value="[integer]"/><!-- 0..1 End position of the window on the starting sequence --> <orientation value="[code]"/><!-- 0..1 sense | antisense --> <strand value="[code]"/><!-- 0..1 watson | crick --> </startingSequence> <edit> <!-- 0..* Changes in sequence from the starting sequence --> <start value="[integer]"/><!-- 0..1 Start position of the edit on the starting sequence --> <end value="[integer]"/><!-- 0..1 End position of the edit on the starting sequence --> <replacementSequence value="[string]"/><!-- 0..1 Allele that was observed --> <replacedSequence value="[string]"/><!-- 0..1 Allele in the starting sequence --> </edit> </relative> </MolecularSequence>
JSON Template
{ "resourceType" : "MolecularSequence",
// from Resource: id, meta, implicitRules, and language
// from DomainResource: text, contained, extension, and modifierExtension
"identifier" : [{ Identifier }], // Unique ID for this particular sequence
"type" : "<code>", // aa | dna | rna
"subject" : { Reference(BiologicallyDerivedProduct|Device|Group|Location|
Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure|
Substance) }, // Subject this sequence is associated too
"specimen" : { Reference(Specimen) }, // Specimen used for sequencing
"device" : { Reference(Device) }, // The method for sequencing
"performer" : { Reference(Organization) }, // Who should be responsible for test result
"literal" : "<string>", // Sequence that was observed
"formatted" : [{ Attachment }], // Embedded file or a link (URL) which contains content to represent the sequence
"relative" : [{ // A sequence defined relative to another sequence
"coordinateSystem" : { CodeableConcept }, // R! Ways of identifying nucleotides or amino acids within a sequence
"resourceType" : "MolecularSequence",
// from Resource: id, meta, implicitRules, and language
// from DomainResource: text, contained, extension, and modifierExtension
"identifier" : [{ Identifier }], // Unique ID for this particular sequence
"type" : "<code>", // aa | dna | rna
"subject" : { Reference(BiologicallyDerivedProduct|Device|Group|Location|
Medication|NutritionProduct|Organization|Patient|Practitioner|Procedure|
Substance) }, // Subject this sequence is associated too
"specimen" : { Reference(Specimen) }, // Specimen used for sequencing
"device" : { Reference(Device) }, // The method for sequencing
"performer" : { Reference(Organization) }, // Who should be responsible for test result
"literal" : "<string>", // Sequence that was observed
"formatted" : [{ Attachment }], // Embedded file or a link (URL) which contains content to represent the sequence
"relative" : [{ // A sequence defined relative to another sequence
"coordinateSystem" : { CodeableConcept }, // R! Ways of identifying nucleotides or amino acids within a sequence  "ordinalPosition" : <integer>, // Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together
"sequenceRange" : { Range }, // Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together
"startingSequence" : { // A sequence used as starting sequence
"genomeAssembly" : { CodeableConcept }, // The genome assembly used for starting sequence, e.g. GRCh38
"ordinalPosition" : <integer>, // Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together
"sequenceRange" : { Range }, // Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together
"startingSequence" : { // A sequence used as starting sequence
"genomeAssembly" : { CodeableConcept }, // The genome assembly used for starting sequence, e.g. GRCh38  "chromosome" : { CodeableConcept }, // Chromosome Identifier
"chromosome" : { CodeableConcept }, // Chromosome Identifier  // sequence[x]: The reference sequence that represents the starting sequence. One of these 3:
"sequenceCodeableConcept" : { CodeableConcept },
"sequenceString" : "<string>",
"sequenceReference" : { Reference(MolecularSequence) },
"windowStart" : <integer>, // Start position of the window on the starting sequence
"windowEnd" : <integer>, // End position of the window on the starting sequence
"orientation" : "<code>", // sense | antisense
"strand" : "<code>" // watson | crick
},
"edit" : [{ // Changes in sequence from the starting sequence
"start" : <integer>, // Start position of the edit on the starting sequence
"end" : <integer>, // End position of the edit on the starting sequence
"replacementSequence" : "<string>", // Allele that was observed
"replacedSequence" : "<string>" // Allele in the starting sequence
}]
}]
}
// sequence[x]: The reference sequence that represents the starting sequence. One of these 3:
"sequenceCodeableConcept" : { CodeableConcept },
"sequenceString" : "<string>",
"sequenceReference" : { Reference(MolecularSequence) },
"windowStart" : <integer>, // Start position of the window on the starting sequence
"windowEnd" : <integer>, // End position of the window on the starting sequence
"orientation" : "<code>", // sense | antisense
"strand" : "<code>" // watson | crick
},
"edit" : [{ // Changes in sequence from the starting sequence
"start" : <integer>, // Start position of the edit on the starting sequence
"end" : <integer>, // End position of the edit on the starting sequence
"replacementSequence" : "<string>", // Allele that was observed
"replacedSequence" : "<string>" // Allele in the starting sequence
}]
}]
}
Turtle Template
@prefix fhir: <http://hl7.org/fhir/> .[ a fhir:MolecularSequence; fhir:nodeRole fhir:treeRoot; # if this is the parser root # from Resource: .id, .meta, .implicitRules, and .language # from DomainResource: .text, .contained, .extension, and .modifierExtension fhir:MolecularSequence.identifier [ Identifier ], ... ; # 0..* Unique ID for this particular sequence fhir:MolecularSequence.type [ code ]; # 0..1 aa | dna | rna fhir:MolecularSequence.subject [ Reference(BiologicallyDerivedProduct|Device|Group|Location|Medication|NutritionProduct| Organization|Patient|Practitioner|Procedure|Substance) ]; # 0..1 Subject this sequence is associated too fhir:MolecularSequence.specimen [ Reference(Specimen) ]; # 0..1 Specimen used for sequencing fhir:MolecularSequence.device [ Reference(Device) ]; # 0..1 The method for sequencing fhir:MolecularSequence.performer [ Reference(Organization) ]; # 0..1 Who should be responsible for test result fhir:MolecularSequence.literal [ string ]; # 0..1 Sequence that was observed fhir:MolecularSequence.formatted [ Attachment ], ... ; # 0..* Embedded file or a link (URL) which contains content to represent the sequence fhir:MolecularSequence.relative [ # 0..* A sequence defined relative to another sequence fhir:MolecularSequence.relative.coordinateSystem [ CodeableConcept ]; # 1..1 Ways of identifying nucleotides or amino acids within a sequence fhir:MolecularSequence.relative.ordinalPosition [ integer ]; # 0..1 Indicates the order in which the sequence should be considered when putting multiple 'relative' elements together fhir:MolecularSequence.relative.sequenceRange [ Range ]; # 0..1 Indicates the nucleotide range in the composed sequence when multiple 'relative' elements are used together fhir:MolecularSequence.relative.startingSequence [ # 0..1 A sequence used as starting sequence fhir:MolecularSequence.relative.startingSequence.genomeAssembly [ CodeableConcept ]; # 0..1 The genome assembly used for starting sequence, e.g. GRCh38 fhir:MolecularSequence.relative.startingSequence.chromosome [ CodeableConcept ]; # 0..1 Chromosome Identifier # MolecularSequence.relative.startingSequence.sequence[x] : 0..1 The reference sequence that represents the starting sequence. One of these 3 fhir:MolecularSequence.relative.startingSequence.sequenceCodeableConcept [ CodeableConcept ] fhir:MolecularSequence.relative.startingSequence.sequenceString [ string ] fhir:MolecularSequence.relative.startingSequence.sequenceReference [ Reference(MolecularSequence) ] fhir:MolecularSequence.relative.startingSequence.windowStart [ integer ]; # 0..1 Start position of the window on the starting sequence fhir:MolecularSequence.relative.startingSequence.windowEnd [ integer ]; # 0..1 End position of the window on the starting sequence fhir:MolecularSequence.relative.startingSequence.orientation [ code ]; # 0..1 sense | antisense fhir:MolecularSequence.relative.startingSequence.strand [ code ]; # 0..1 watson | crick ]; fhir:MolecularSequence.relative.edit [ # 0..* Changes in sequence from the starting sequence fhir:MolecularSequence.relative.edit.start [ integer ]; # 0..1 Start position of the edit on the starting sequence fhir:MolecularSequence.relative.edit.end [ integer ]; # 0..1 End position of the edit on the starting sequence fhir:MolecularSequence.relative.edit.replacementSequence [ string ]; # 0..1 Allele that was observed fhir:MolecularSequence.relative.edit.replacedSequence [ string ]; # 0..1 Allele in the starting sequence ], ...; ], ...; ]
Changes since Release 4
| MolecularSequence | |
| MolecularSequence.subject |
|
| MolecularSequence.literal |
|
| MolecularSequence.formatted |
|
| MolecularSequence.relative |
|
| MolecularSequence.relative.coordinateSystem |
|
| MolecularSequence.relative.ordinalPosition |
|
| MolecularSequence.relative.sequenceRange |
|
| MolecularSequence.relative.startingSequence |
|
| MolecularSequence.relative.startingSequence.genomeAssembly |
|
| MolecularSequence.relative.startingSequence.chromosome |
|
| MolecularSequence.relative.startingSequence.sequence[x] |
|
| MolecularSequence.relative.startingSequence.windowStart |
|
| MolecularSequence.relative.startingSequence.windowEnd |
|
| MolecularSequence.relative.startingSequence.orientation |
|
| MolecularSequence.relative.startingSequence.strand |
|
| MolecularSequence.relative.edit |
|
| MolecularSequence.relative.edit.start |
|
| MolecularSequence.relative.edit.end |
|
| MolecularSequence.relative.edit.replacementSequence |
|
| MolecularSequence.relative.edit.replacedSequence |
|
| MolecularSequence.coordinateSystem |
|
| MolecularSequence.patient |
|
| MolecularSequence.quantity |
|
| MolecularSequence.referenceSeq |
|
| MolecularSequence.variant |
|
| MolecularSequence.observedSeq |
|
| MolecularSequence.quality |
|
| MolecularSequence.readCoverage |
|
| MolecularSequence.repository |
|
| MolecularSequence.pointer |
|
| MolecularSequence.structureVariant |
|
See the Full Difference for further information
Additional definitions: Master Definition XML + JSON, XML Schema/Schematron + JSON Schema, ShEx (for Turtle) + see the extensions, the spreadsheet version & the dependency analysis
| Path | Definition | Type | Reference |
|---|---|---|---|
| MolecularSequence.type | Type if a sequence -- DNA, RNA, or amino acid sequence. | Required | sequenceType |
| MolecularSequence.relative.coordinateSystem | Extensible | http://loinc.org/LL5323-2/  | |
| MolecularSequence.relative.startingSequence.genomeAssembly | Extensible | http://loinc.org/LL1040-6/  | |
| MolecularSequence.relative.startingSequence.chromosome | Required | http://loinc.org/LL2938-0/  | |
| MolecularSequence.relative.startingSequence.sequence[x] | Example | ?? | |
| MolecularSequence.relative.startingSequence.orientation | Type for orientation. | Required | orientationType |
| MolecularSequence.relative.startingSequence.strand | Type for strand. | Required | strandType |
| UniqueKey | Level | Location | Description | Expression |
 msq-5 msq-5 | Rule | MolecularSequence.relative.startingSequence | Both genomeAssembly and chromosome must be both contained if either one of them is contained | (chromosome.empty() and genomeAssembly.empty()) or (chromosome.exists() and genomeAssembly.exists()) |
 msq-6 msq-6 | Rule | MolecularSequence.relative.startingSequence | Have and only have one of the following elements in startingSequence: 1. genomeAssembly; 2 sequence | (genomeAssembly.count()+sequenceCodeableConcept.count()+ sequenceReference.count()+ sequenceString.count()) = 1 |
This resource supports three patterns for representing a sequence of interest:
The MolecularSequence resource is designed to represent a single sequence in an instance. Each sequence might have multiple representations, but implementers SHALL ensure all representations are for the same sequence.
MolecularSequence.literal: This string element can be used to hold the sequence as a string of characters.
MolecularSequence.formatted: This Attachment is used to refer to the sequence as embedded file content or via a URL reference.
This method can be used to refer to sequence data from an an external source. If the sequence is referring to a GA4GH repository, the MolecularSequence.formatted.url should refer to a GA4GH compliant endpoint that conforms to GA4GH data models.
MolecularSequence.relative: This complex element is used for encoding sequence. When the information of starting sequence and edits are provided, the observed sequence will be derived. Here is a picture below:
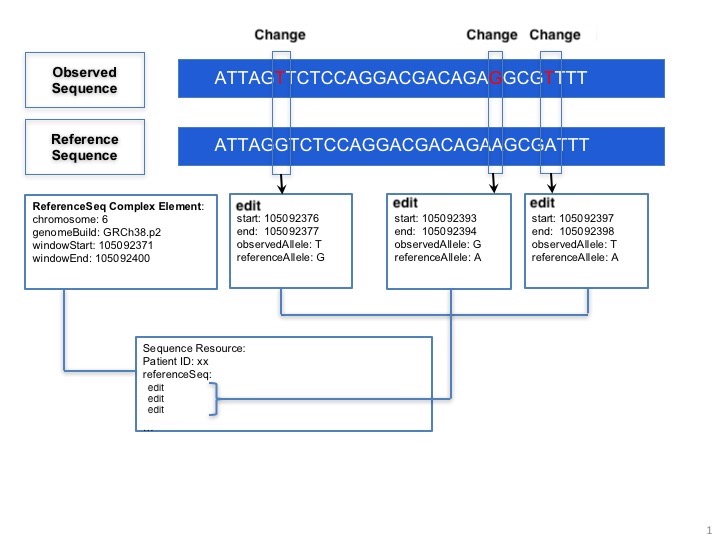
MolecularSequence.relative.ordinalPosition: Indicates the order in which the sequence should be considered when putting multiple relative instances together.
MolecularSequence.relative.sequenceRange: Indicates the nucleotide range in the composed sequence when multiple relative instances are used together.
These attributes help to clarify what sequence is being representing with less computation/inference on the recipient side. Implementers SHOULD use sequenceRange first to determine order as the most reliable. If sequenceRange is not present then ordinalPosition SHOULD be used. Finally, if both sequenceRange and ordinalPosition are absent, then the order of the relative data elements SHOULD be used to calculate a composition. It is the responsibility of the data sender to ensure the message can be consistently understood. Additionally, gaps in sequenceRange are considered intentional (i.e. the composed sequence contains a sequence of N's, the placeholder nucleotide, for the gap range).
In a FGFR2:MET Fusion use case, where the fusion was uncovered through RNA sequencing, a partial representation can be found here.
MolecularSequence.relative.startingSequence: There are four optional ways to represent a starting sequence in MolecularSequence resource:
MolecularSequence.relative.startingSequence.sequenceCodeableConcept: Starting sequence id in public database;MolecularSequence.relative.startingSequence.sequenceString: Starting sequence string; MolecularSequence.relative.startingSequence.sequenceReference: Reference to starting sequence stored in another sequence entity; MolecularSequence.relative.startingSequence.genomeAssembly, MolecularSequence.relative.startingSequence.chromosome: The combination of genome assembly and chromosome.
The MolecularSequence.relative.startingSequence.windowStart and MolecularSequence.relative.startingSequence.windowEnddefines a range from the starting sequence that is used to define a subsequence used as the starting sequence.
When saving the sequence information, the nucleic acid will be numbered with order. Some representations use a 0-based system (e.g. GA4GH API, BAM files) while some use a 1-based system (e.g. VCF file format). The element coordinateSystem contains this information.
MolecularSequence.relative.coordinateSystem binds to a LOINC answer list, please review those answers here  as well as the detailed description found here
as well as the detailed description found here  .
.
Here are two examples:
There are lots of definition concerning with the directionality of DNA or RNA. Here we are using MolecularSequence.relative.startingSequence.orientation and MolecularSequence.relative.startingSequence.strand. Orientation represents the sense of the sequence, which has different meanings depending on the MolecularSequence.type. Strand represents the sequence writing order. Watson strand refers to 5' to 3' top strand (5' -> 3'), whereas Crick strand refers to 5' to 3' bottom strand (3' <- 5').
Only two possible values can be made by strand, watson and crick. Since the directionality of the sequence string might be represented in different word in different omics scenario, below are simple example of how to map other expressions into its correlated value:
| Watson | Crick |
|---|---|
| 5′-to-3′ direction | 3′-to-5′ direction |
| +1 | -1 |
| Sense | Antisense |
| Positive | Negative |
There are attributes where the sequence is represented as a string of characters.
relative.startingSequence.sequenceStringrelative.edit.replacementSequencerelative.edit.replacedSequenceliteral
The characters used in these string representations of a sequence should be constrained to the IUPAC codes found here https://www.bioinformatics.org/sms2/iupac.html  .
.
Search parameters for this resource. The common parameters also apply. See Searching for more information about searching in REST, messaging, and services.
| Name | Type | Description | Expression | In Common |
| identifier | token | The unique identity for a particular sequence | MolecularSequence.identifier | |
| patient | reference | The subject that the sequence is about | MolecularSequence.subject (Practitioner, Group, Organization, BiologicallyDerivedProduct, NutritionProduct, Device, Medication, Patient, Procedure, Substance, Location) | |
| subject | reference | The subject that the sequence is about | MolecularSequence.subject (Practitioner, Group, Organization, BiologicallyDerivedProduct, NutritionProduct, Device, Medication, Patient, Procedure, Substance, Location) | |
| type | token | Amino Acid Sequence/ DNA Sequence / RNA Sequence | MolecularSequence.type |