This page is part of the Genetic Reporting Implementation Guide (v3.0.0-ballot: STU 3 Ballot 1) based on FHIR (HL7® FHIR® Standard) R4. The current version which supersedes this version is 3.0.0. For a full list of available versions, see the Directory of published versions
| Official URL: http://hl7.org/fhir/uv/genomics-reporting/OperationDefinition/find-population-molecular-consequences | Version: 3.0.0-ballot | |||
| Active as of 2023-12-18 | Computable Name: FindPopulationMolecularConsequences | |||
Retrieve count or list of patients having molecular consequences.
Retrieve count or list of patients having molecular consequences. More specifically, this operation retrieves the count +/- list of patients that have molecular consequences involving specific featureConsequences, derived from specific variants.
A patient meets numerator criteria if they have at least one molecular consequence matching the query parameters.
As shown in the following picture, this operation can return previously instantiated implications and/or dynamically computed implications. Specific implementations can indicate their capabilities using a FHIR Capability Statement. Rules around the retention of dynamically computed implications are outside the scope of this operation, but a server could potentially instantiate those results based on the Therapeutic Implication, Diagnostic Implication, or Molecular Consequence FHIR profiles.
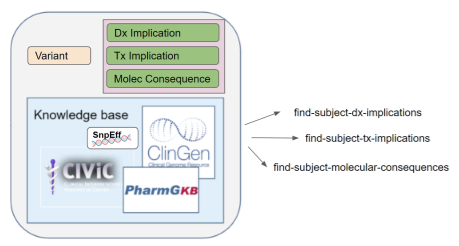
Population queries are designed to return a count of patients that match each item sought, with or without a list of patients matching the item(s) sought.
Parameters
| Use | Name | Scope | Cardinality | Type | Binding | Documentation |
| IN | variants | 0..* | string (string) | List of variants from which implications are derived. Must be in HGVS or SPDI format. | ||
| IN | featureConsequences | 0..* | string (token) | List of consequences sought. Must be in token or codesystem|code format. (These will generally be coded with Sequence Ontology codes under SO:0001537) | ||
| IN | genomicSourceClass | 0..1 | string (token) | Enables an App to limit results to those that are 'germline' or 'somatic'. Default is to include variants irrespective of genomic source class. | ||
| IN | includePatientList | 0..1 | boolean | Include list of matching patients if set to true. Default=false. | ||
| OUT | consequences | 1..1 | ||||
| OUT | consequences.numerator | 1..1 | Quantity | Count of patients meeting criteria | ||
| OUT | consequences.denominator | 0..1 | Quantity | Count of patients in the cohort searched | ||
| OUT | consequences.subject | 0..* | string | Patient ID. Include if includePatientList=true |
Contents:
Valid response codes are shown in the following table. Additional response codes (e.g. 5xx server error) may also be encountered.
| Response Code | Description |
|---|---|
| 200 | Successfully executed request |
| 400 | ERROR: Invalid query parameters |
| 404 | ERROR: Patient not found |
| 422 | ERROR: Failed LiftOver |
As part of a quality assurance program, an oncologist seeks to identify patients harboring putative splice site variants in order to further assess disparities in molecularly guided treatment.